Virus populations that attack human gut bacteria change rapidly
31 July 2013
The viral population that preys on bacteria in the human gut can undergo rapid change over short time periods, according to a new genetic study by researchers from the Perelman School of Medicine at the University of Pennsylvania. The evolution and variety of this 'virome' can affect susceptibility and resistance to disease among individuals, and also the effectiveness of drugs.
The research team, led by Professor of Microbiology Dr Frederic Bushman, extracted viral particles from human stool samples using several methods over 884 days. They then isolated and analyzed DNA contigs (contiguous sequences) using ultra-deep genome sequencing. The result was the longest, most extensive picture of the workings of the human virome yet obtained, according to Prof Bushman. Their work has been published in the Proceedings of the National Academy of Sciences.
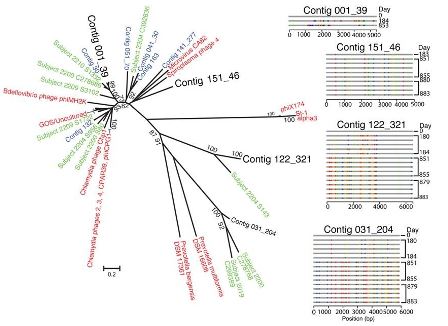
Phylogenetic tree of microphages detected in
this and other studies. The four microphage contigs with the
highest substitution rates observed in the PNAS study are shown in
large black lettering. The scale bar indicates the proportion of
amino acid substitutions within the 919 amino acid major coat
protein, which was aligned to make the tree.
“Bacterial viruses are predators on bacteria, so they mold their populations,” said Bushman. “Bacterial viruses also transport genes for toxins, virulence factors that modify the phenotype of their bacterial host.” In this way, an innocent, benign bacterium living inside the body can be transformed by an invading virus into a dangerous threat.
The researchers found that while approximately 80% of the viral types identified remained mostly unchanged over the course of the study, certain viral species changed so substantially over time that, as Bushman notes, “You could say we observed speciation events.”
This was particularly true in the Microviridae group, which are bacteriophages with single-stranded circular DNA genomes. Several genetic mechanisms drove the changes, including substitution of base chemicals; diversity-generating retroelements, in which reverse transcriptase enzymes introduce mutations into the genome; and CRISPRs (Clustered Regularly Interspaced Short Palindromic Repeats), in which pieces of the DNA sequences of bacteriophages are incorporated as spacers in the genomes of bacteria.
Such rapid evolution of the virome was perhaps the most surprising finding for the research team. Bushman notes that “different people have quite different bacteria in their guts, so the viral predators on those bacteria are also different. However, another reason people are so different from each other in terms of their virome, emphasized in this paper, is that some of the viruses, once inside a person, are changing really fast. So some of the viral community diversifies and becomes unique within each individual.”
Since humans acquire the bacterial population — and its accompanying virome — after birth from food and other environmental factors, it’s logical that the microbial population living within each of us would differ from person to person. But this work, say the researchers, demonstrates that another major explanatory factor is the constant evolution of the virome within the body. That fact has important implications for the ways in which susceptibility and resistance to disease can differ among individuals, as well as the effectiveness of various drugs and other treatments.