Stucture of AIDS-like virus enzyme solved in three weeks by online gamers
23 September 2011
The structure of a retrovirus enzyme whose configuration had stumped scientists for more than a decade has been solved in three weeks by gamers. The gamers achieved their discovery by playing Foldit, an online game developed by the University of Washington that allows players to collaborate and compete in predicting the structure of protein molecules.
After scientists repeatedly failed to piece together the structure of a protein-cutting enzyme from an AIDS-like virus, they called in the Foldit players. The scientists challenged the gamers to produce an accurate model of the enzyme. They did it in only three weeks.
This class of enzymes, called retroviral proteases, has a critical role in how the AIDS virus matures and proliferates. Intensive research is under way to try to find anti-AIDS drugs that can block these enzymes, but efforts were hampered by not knowing exactly what the retroviral protease molecule looks like.
"We wanted to see if human intuition could succeed where automated methods had failed," said Dr Firas Khatib of the University of Washington Department of Biochemistry. Khatib is a researcher in the protein structure lab of Dr David Baker, professor of biochemistry.
Remarkably, the gamers generated models good enough for the researchers to refine and, within a few days, determine the enzyme's structure. Equally amazing, surfaces on the molecule stood out as likely targets for drugs to de-active the enzyme.
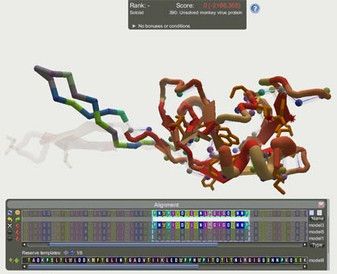
A screenshot of the "Unsolved monkey virus
protein" Foldit puzzle,
highlighting the Alignment Tool used by
Foldit players to
combine regions from multiple models into a
single hybrid structure.
"These features provide exciting opportunities for the design of retroviral drugs, including AIDS drugs," wrote the authors of a paper appearing Sept. 18 in Nature Structural & Molecular Biology. The scientists and gamers are listed as co-authors.
This is the first instance that the researchers are aware of in which gamers solved a longstanding scientific problem.
Fold-it was created by computer scientists at the University of Washington Center for Game Science in collaboration with the Baker lab.
Direct manipulation tools, as well as assistance from a computer program called Rosetta, encourage participants to configure graphics into a workable protein model. Teams send in their answers, and UW researchers constantly improve the design of the game and its puzzles by analyzing the players' problem-solving strategies.
Figuring out the shape and misshape of proteins contributes to research on causes of and cures for cancer, Alzheimer's, immune deficiencies and a host of other disorders, as well as to environmental work on biofuels.
Dr. Seth Cooper, of the UW Department of Computing Science and Engineering, is a co-creator of Foldit and its lead designer and developer. He studies human-computer exploration methods and the co-evolution of games and players.
"People have spatial reasoning skills, something computers are not yet good at," Cooper said. "Games provide a framework for bringing together the strengths of computers and humans. The results in this week's paper show that gaming, science and computation can be combined to make advances that were not possible before."
Games like Foldit are evolving. To piece together the retrovirus enzyme structure, Cooper said, gamers used a new Alignment Tool for the first time to copy parts of know molecules and test their fit in an incomplete model.
"The ingenuity of game players," Khatib said, "is a formidable force that, if properly directed, can be used to solve a wide range of scientific problems.
According to Popovic, "Foldit shows that a game can turn novices into domain experts capable of producing first-class scientific discoveries. We are currently applying the same approach to change the way math and science are taught in school."
More information
1. Khatib F et al. Crystal structure of a monomeric retroviral protease solved by protein folding game players. Nature Structural and Molecular Biology. Published online 18 September 2011; doi:10.1038/nsmb.2119
2. Play Foldit: http://fold.it/portal/
3. Video on Foldit: