First 3-D image of antibody gene
29 April 2008
Scientists at the University of California, San Diego have shown for the first time how a genome is organized in three-dimensional space. They used a multidisciplinary mix of geometry, biological research and techniques developed to solve problems on supercomputers.
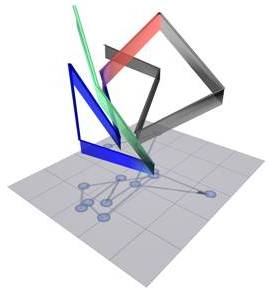 |
| The 3-D structure of the immunoglobulin locus in B cells, showing the relative positions of the different portions of the immunoglobulin genes. Grey objects indicate constant regions. Blue objects indicate proximal variable regions. Green objects indicate distal variable regions. Red line indicates the linker connecting the proximal variable and joining regions. |
Researchers led by Cornelis Murre, a professor of biology at UC San Diego, and Steve Cutchin, senior scientist for visualization services at the San Diego Supercomputer Center (SDSC), used the gene encoding the immunoglobulin heavy chain locus — responsible for generating diverse kinds of antibodies — to demonstrate the structure of the genome. The findings are published in the April issue of Cell [1].
The observations, the researchers say, permit an insight into the structure of the human genome, which until now has remained elusive.
Because the genome is the most essential part of the cell for storing and accessing genetic information, the complete DNA sequence of a wide variety of genomes has been revealed in studies performed in a large number of laboratories — “a tremendous success that has provided insight into mechanisms that underpin the development of a wide variety of diseases,” the authors say.
However, Murre said, “it has remained unclear as to how the genome is organized in three-dimensional space. This is an important issue since the regulation of gene expression is controlled by interactions of genomic elements that are separated by large genomic distances. Thus, our team wanted to determine how the genome is structured within the nucleus.”
The experiments described in the Cell paper, he said, provide a first glimpse into this question. “As a model system, we used the gene encoding for the immunoglobulin heavy chain locus, because it is responsible for generating the wide diversity of antibodies.”
Having measured the distances that separate the various parts of the gene, Murre said, the researchers, in collaboration with Cutchin at the SDSC, then used geometry to resolve the first structure of a genetic locus.
His work, said Cutchin, involved computational geometry, scientific visualization, computational methods and numerical methods. “The resulting structure shows that the antibody gene is organized into ‘flower-like’ structures that are connected by linkers,” said Murre. “These flowers contain the various parts that ultimately generate the wide variety of antibodies. This is the first time that geometry has been used to determine the structure of a genetic locus. Ultimately, the same approach should be used to elucidate the structure of the entire human genome.”
Reference1. Suchit Jhunjhunwala, Menno C van Zelm, Mandy M Peak, Steve Cutchin,
Roy Riblet, Jacques JM van Dongen, Frank G Grosveld, Tobias A Knoch, and
Cornelis Murre. The 3-D Structure of the Immunoglobulin Heavy Chain
Locus: Implications for Long-Range Genomic Interactions. Cell, 18
April 2008, p 265.
DOI:
http://dx.doi.org/10.1016/j.cell.2008.03.024